calculate_summary_distances_of_cells_to_borders
Source:R/calculate_summary_distances_of_cells_to_borders.R
calculate_summary_distances_of_cells_to_borders.RdReturns the mean, median and standard deviation of the distances between a specified cell type to the border.
Usage
calculate_summary_distances_of_cells_to_borders(
sce_object,
cell_types_of_interest,
feature_colname = "Cell.Type"
)Arguments
- sce_object
SingleCellExperiment object whose metadata contains the information of tumour structure and cell distances to tumour border (has column "Region" and "Distance.To.Border").
- cell_types_of_interest
String Vector of cell types to consider.
- feature_colname
String specifying which column the interested cell types are from.
Examples
sce_border <- identify_bordering_cells(SPIAT::defined_image,
reference_cell = "Tumour", feature_colname = "Cell.Type", n_to_exclude = 10)
#> [1] "The alpha of Polygon is: 63.24375"
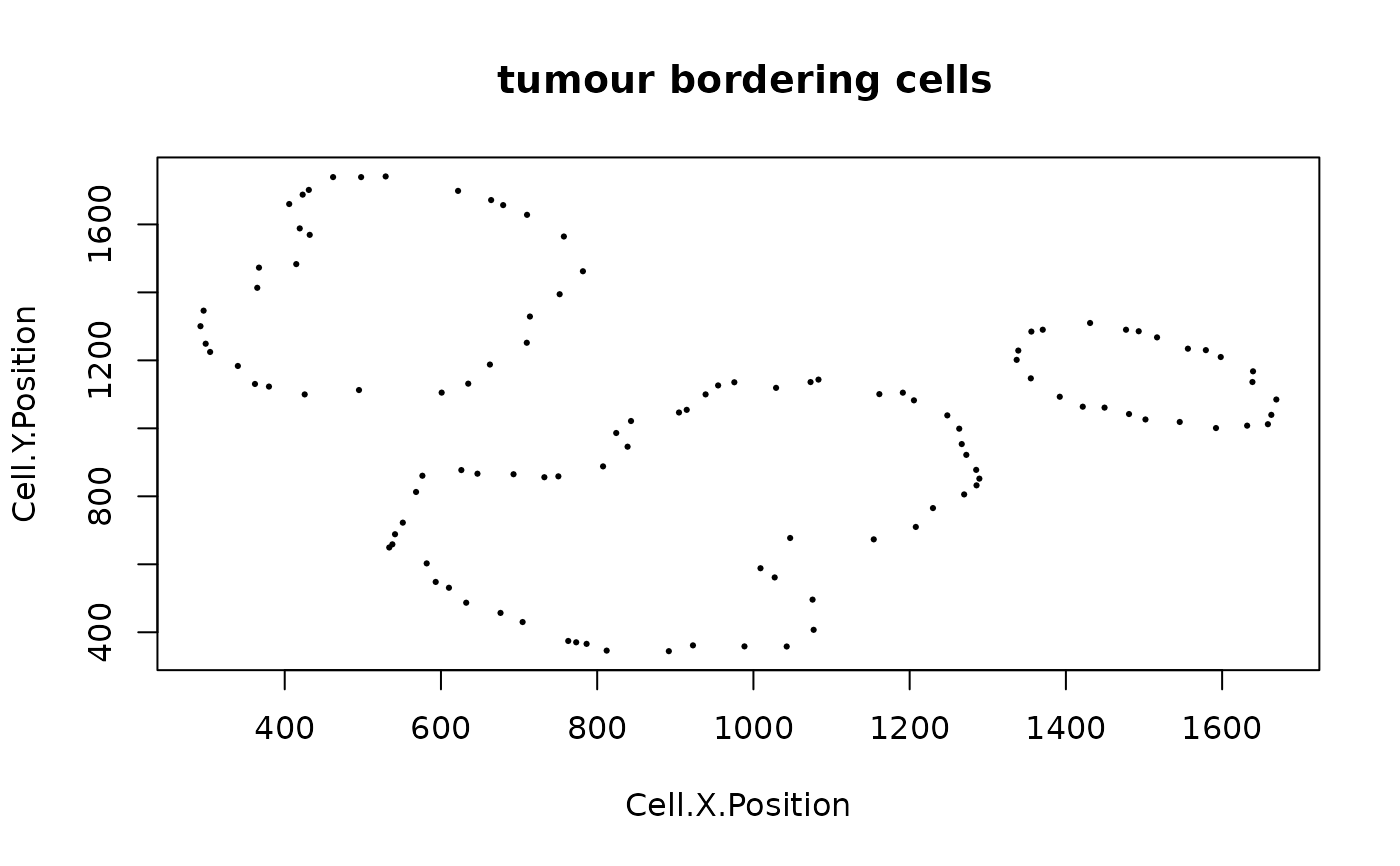 sce_dist <- calculate_distance_to_tumour_margin(sce_border)
#> [1] "Markers had been selected in pair-wise distance calculation: "
#> [1] "Non-border" "Border"
sce_structure <- define_structure(sce_dist, names_of_immune_cells =
c("Immune1","Immune2","Immune3"), feature_colname = "Cell.Type",
n_margin_layers = 5)
calculate_summary_distances_of_cells_to_borders(sce_structure,
cell_types_of_interest = c("Immune1","Immune3"),feature_colname = "Cell.Type")
#> Cell.Type Area Min_d Max_d Mean_d Median_d
#> 1 All_cell_types_of_interest Tumor_area 10.93225 192.4094 86.20042 88.23299
#> 2 All_cell_types_of_interest Stroma 10.02387 971.5638 195.10637 101.95113
#> 3 Immune1 Tumor_area Inf -Inf NaN NA
#> 4 Immune1 Stroma 84.20018 970.7749 346.14096 301.01535
#> 5 Immune3 Tumor_area 10.93225 192.4094 86.20042 88.23299
#> 6 Immune3 Stroma 10.02387 971.5638 102.79227 68.19218
#> St.dev_d
#> 1 45.27414
#> 2 194.68507
#> 3 NA
#> 4 187.04247
#> 5 45.27414
#> 6 131.32714
sce_dist <- calculate_distance_to_tumour_margin(sce_border)
#> [1] "Markers had been selected in pair-wise distance calculation: "
#> [1] "Non-border" "Border"
sce_structure <- define_structure(sce_dist, names_of_immune_cells =
c("Immune1","Immune2","Immune3"), feature_colname = "Cell.Type",
n_margin_layers = 5)
calculate_summary_distances_of_cells_to_borders(sce_structure,
cell_types_of_interest = c("Immune1","Immune3"),feature_colname = "Cell.Type")
#> Cell.Type Area Min_d Max_d Mean_d Median_d
#> 1 All_cell_types_of_interest Tumor_area 10.93225 192.4094 86.20042 88.23299
#> 2 All_cell_types_of_interest Stroma 10.02387 971.5638 195.10637 101.95113
#> 3 Immune1 Tumor_area Inf -Inf NaN NA
#> 4 Immune1 Stroma 84.20018 970.7749 346.14096 301.01535
#> 5 Immune3 Tumor_area 10.93225 192.4094 86.20042 88.23299
#> 6 Immune3 Stroma 10.02387 971.5638 102.79227 68.19218
#> St.dev_d
#> 1 45.27414
#> 2 194.68507
#> 3 NA
#> 4 187.04247
#> 5 45.27414
#> 6 131.32714